This permutation works with the example matrix in In : B = np.matrix('1 0 0 0 1 0 0 0 0 1 1 0 0 1 0 1 0 1 1 0 1 0 0 0 0 0 0 The MATLAB docs say that with this permutation, the eigenvalues remain the same. In numpy terms, that means: In : I,J=np.ix_(permutation_array,permutation_array)Īnd indeed there are more 0 bands in the 2 off diagonal corners.Īnd using the bandwidth calculation on the MATLAB page, In : i,j=A.nonzero() It will probably have to be a generator (of some sort) so I can test each permutation as it is generated otherwise it will eat up the memory in no time. This is a permutation r such that S(r,r) tends to have its nonzero elements closer to the diagonal. I need it to be resume-able from the nth permutation, as there are going to be a lot of permutations I cant test them all in one run. R = symrcm(S) returns the symmetric reverse Cuthill-McKee ordering of S. I don't see any improvement in clustering of 1s in res. 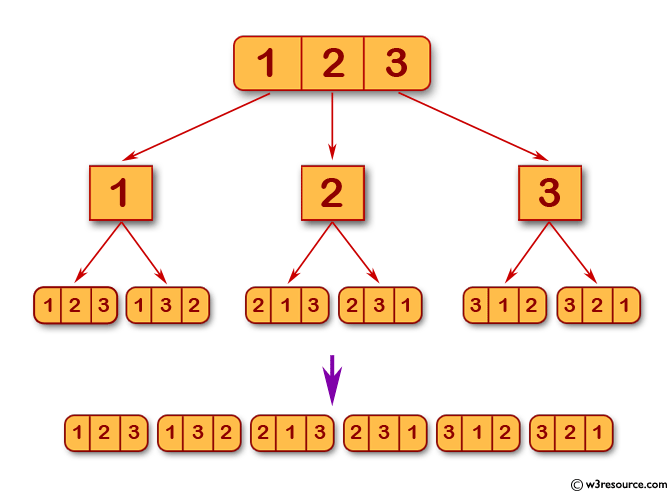 
You on the other hand appear to be doing: In : res = sparse.lil_matrix(A.shape,dtype=A.dtype) I see some clustering maybe the best we can expect from a random matrix. My first inclination is to use such an array to simply index rows of the original matrix: In : A.A (oops, used the wrong matrix here): In : permutation_array = csgraph.reverse_cuthill_mckee(A, False) In : A = sparse.csr_matrix((np.ones().astype(np.uint8),ran.indices, ran.indptr)) Make a random matrix (float) and convert it to a uint8 (beware, csr calculations might not work with this dtype): In : ran=sparse.random(10,10.3, format='csr') I wonder if you are applying the permutation array correctly. So this array is something like: (size = 200,000) The result of the reverse_cuthill_mckee call is an array which is like a tupel containing the indices for my permutation. Permutation_array = csgraph.reverse_cuthill_mckee(A, false) 
I tried a for-loop but my matrix has dimension like 200,000 x 150,000 and it takes too much time. Now is there any efficient solution for doing this permutation on my sparse csr_matrix in any other sparse matrix (csr, lil_matrix, etc)? The result of this method is a permutation array whichs gives me the indices of how to permutate the rows of my matrix as I understood. "Random selection from itertools.I used the Scipy Reverse Cuthill-McKee implementation ( _cuthill_mckee) for creating a band matrix using a (high-dimensional) sparse csr_matrix. See also more_itertools docs for further information on this tool.įor those interested, here is the actual recipe.įrom the itertools recipes: def random_permutation(iterable, r=None): random_permute_generator(iterable, n=5000). List(random_permute_generator(range(10), n=20))įor your specific problem, substitute the iterable and number of calls n with the appropriate values, e.g. """Yield a random permuation of an iterable n times.""" We will implement this generator and demonstrate random results with an abridged example: def random_permute_generator(iterable, n=10): We can make a generator that yields these results for n calls. For convenience I use a third-party library, more_itertools, that implements this recipe for us: import more_itertools as mit You can try implementing the random_permutation itertools recipes. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
